Temperature as an Extra Dimension in Multidimensional Protein NMR Spectroscopy
Dr. Alexandra Shchukina, Paweł Małecki, Dr. Borja Mateos, Dr. Michał Nowakowski, Dr. Mateusz Urbańczyk, Dr. Georg Kontaxis, Dr. Paweł Kasprzak, Clara Conrad‐Billroth, Prof. Robert Konrat, Prof. Krzysztof Kazimierczuk
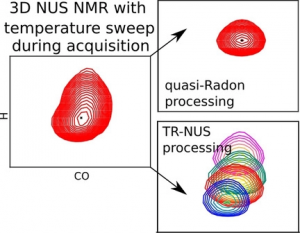
NMR spectroscopy is a particularly informative method for studying protein structures and dynamics in solution; however, it is also one of the most time‐consuming. Modern approaches to biomolecular NMR spectroscopy are based on lengthy multidimensional experiments, the duration of which grows exponentially with the number of dimensions. The experimental time may even be several days in the case of 3D and 4D spectra. Moreover, the experiment often has to be repeated under several different conditions, for example, to measure the temperature‐dependent effects in a spectrum (temperature coefficients (TCs)). Herein, a new approach that involves joint sampling of indirect evolution times and temperature is proposed. This allows TCs to be measured through 3D spectra in even less time than that needed to acquire a single spectrum by using the conventional approach. Two signal processing methods that are complementary, in terms of sensitivity and resolution, 1) dividing data into overlapping subsets followed by compressed sensing reconstruction, and 2) treating the complete data set with a variant of the Radon transform, are proposed. The temperature‐swept 3D HNCO spectra of two intrinsically disordered proteins, osteopontin and CD44 cytoplasmic tail, show that this new approach makes it possible to determine TCs and their non‐linearities effectively. Non‐linearities, which indicate the presence of a compact state, are particularly interesting. The complete package of data acquisition and processing software for this new approach are provided.

